Discover SWOT L3 Unsmoothed data, and compare with CMEMS L4 SLA and IFREMER SST data
Tutorial Objectives
Download Swot data with cycle/passes numbers, using
altimetry_downloader_avisoSelect data intersecting a geographical area, using
altimetry.ioDownload CMEMS map using
copernicusmarineclient, and plot Swot SLA with CMEMS SLADownload SST map using
copernicusmarineclient and plot Swot SLA with SST
Note
Required environment to run this notebook:
xarraynumpymatplotlib+cartopy+cmoceanaltimetry_downloader_aviso: see documentation.altimetry.io: available here.copernicusmarine: see documentation.
Import + code
import warnings
warnings.filterwarnings('ignore')
# Configure logging if you want more details
import logging
logging.basicConfig(level=logging.INFO)
from pathlib import Path
import xarray as xr
import numpy as np
import cartopy.crs as ccrs
import matplotlib.cm as mplcm
import matplotlib.pyplot as plt
import cmocean
import copernicusmarine
def plot_datasets(datasets, variable, vminmax, title, extent=None):
cb_args = dict(
add_colorbar=True,
cbar_kwargs={"shrink": 0.3}
)
plot_kwargs = dict(
x="longitude",
y="latitude",
cmap="Spectral_r",
vmin=vminmax[0],
vmax=vminmax[1],
)
fig, ax = plt.subplots(figsize=(12, 8), subplot_kw=dict(projection=ccrs.PlateCarree()))
if extent: ax.set_extent(extent)
for ds in datasets:
ds[variable].plot.pcolormesh(
ax=ax,
**plot_kwargs,
**cb_args)
cb_args=dict(add_colorbar=False)
ax.set_title(title)
ax.coastlines()
gls = ax.gridlines(draw_labels=True)
gls.top_labels=False
gls.right_labels=False
return ax
Parameters
Define a local filepath to download files
output_dir= Path.home() / "TMP_DATA"
Define cycles and half_orbits numbers to download
Note
To look for Swot passes in a predefined geographical area and temporal period, use Search Swot tool.
cycle_number = [2]
pass_number = [14, 169]
# English channel
bbox = (-7, 46, 4, 52)
localbox = [-7, 4, 46, 52]
Download data using altimetry_downloader_aviso
import altimetry_downloader_aviso as dl_aviso
dl_aviso.get(
'SWOT_L3_LR_SSH_Unsmoothed',
output_dir=output_dir,
cycle_number=cycle_number,
pass_number=pass_number,
)
INFO:altimetry_downloader_aviso.catalog_client.client:Fetching products from Aviso's catalog...
INFO:altimetry_downloader_aviso.catalog_client.granule_discoverer:Filtering SWOT_L3_LR_SSH_Unsmoothed product with filters {'cycle_number': [2], 'pass_number': [14, 169]}...
INFO:altimetry_downloader_aviso.core:2 files to download. 0 files already exist.
INFO:altimetry_downloader_aviso.tds_client:File /home/atonneau/TMP_DATA/SWOT_L3_LR_SSH_Unsmoothed_002_014_20230811T132741_20230811T141908_v2.0.1.nc downloaded.
INFO:altimetry_downloader_aviso.tds_client:File /home/atonneau/TMP_DATA/SWOT_L3_LR_SSH_Unsmoothed_002_169_20230817T022159_20230817T031326_v2.0.1.nc downloaded.
['/home/atonneau/TMP_DATA/SWOT_L3_LR_SSH_Unsmoothed_002_014_20230811T132741_20230811T141908_v2.0.1.nc',
'/home/atonneau/TMP_DATA/SWOT_L3_LR_SSH_Unsmoothed_002_169_20230817T022159_20230817T031326_v2.0.1.nc']
Open data using altimetry.io
from altimetry.io import AltimetryData, FileCollectionSource
Open data source
alti_data = AltimetryData(
source=FileCollectionSource(
path=output_dir,
ftype="SWOT_L3_LR_SSH",
subset="Unsmoothed"
),
)
Query data
ds_unsmoothed_14 = alti_data.query_orbit(
cycle_number=cycle_number,
pass_number=14,
variables=['longitude', 'latitude', 'time', 'ssha_unfiltered', 'ssha_filtered', 'ssha_unedited', 'sigma0', 'quality_flag'],
polygon=bbox
)
INFO:fcollections.implementations.optional._predicates:The bbox intersects with pass numbers (calval phase): [16]
INFO:fcollections.implementations.optional._predicates:The bbox intersects with pass numbers (science phase): [14, 42, 70, 85, 98, 113, 126, 141, 169, 197, 225, 264, 292, 320, 348, 363, 376, 391, 404, 419, 447, 475, 503, 542, 570]
INFO:fcollections.core._readers:Files to read: 1
INFO:fcollections.implementations.optional._area_selectors:Input longitudes does not fit in one of the known 360° intervals. Using [0, 360] (a copy of the input will be made)
INFO:fcollections.implementations.optional._area_selectors:Size of the dataset matching the bbox: {'num_lines': 3100, 'num_pixels': 519}
ds_unsmoothed_169 = alti_data.query_orbit(
cycle_number=cycle_number,
pass_number=169,
variables=['longitude', 'latitude', 'time', 'ssha_unfiltered', 'ssha_filtered', 'ssha_unedited', 'sigma0', 'quality_flag'],
polygon=bbox
)
INFO:fcollections.implementations.optional._predicates:The bbox intersects with pass numbers (calval phase): [16]
INFO:fcollections.implementations.optional._predicates:The bbox intersects with pass numbers (science phase): [14, 42, 70, 85, 98, 113, 126, 141, 169, 197, 225, 264, 292, 320, 348, 363, 376, 391, 404, 419, 447, 475, 503, 542, 570]
INFO:fcollections.core._readers:Files to read: 1
INFO:fcollections.implementations.optional._area_selectors:Input longitudes does not fit in one of the known 360° intervals. Using [0, 360] (a copy of the input will be made)
INFO:fcollections.implementations.optional._area_selectors:Size of the dataset matching the bbox: {'num_lines': 3102, 'num_pixels': 519}
Visualise data
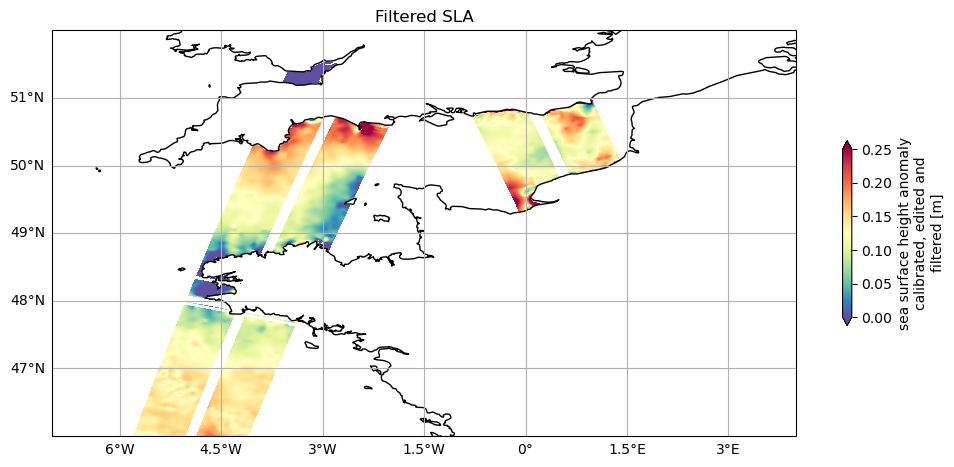
Let’s plot SLA for each cycle
plot_datasets([ds_unsmoothed_169, ds_unsmoothed_14], 'ssha_filtered', (0, 0.25), 'Filtered SLA', extent=localbox)
<GeoAxes: title={'center': 'Filtered SLA'}, xlabel='longitude (degrees East)\n[degrees_east]', ylabel='latitude (positive N, negative\nS) [degrees_north]'>
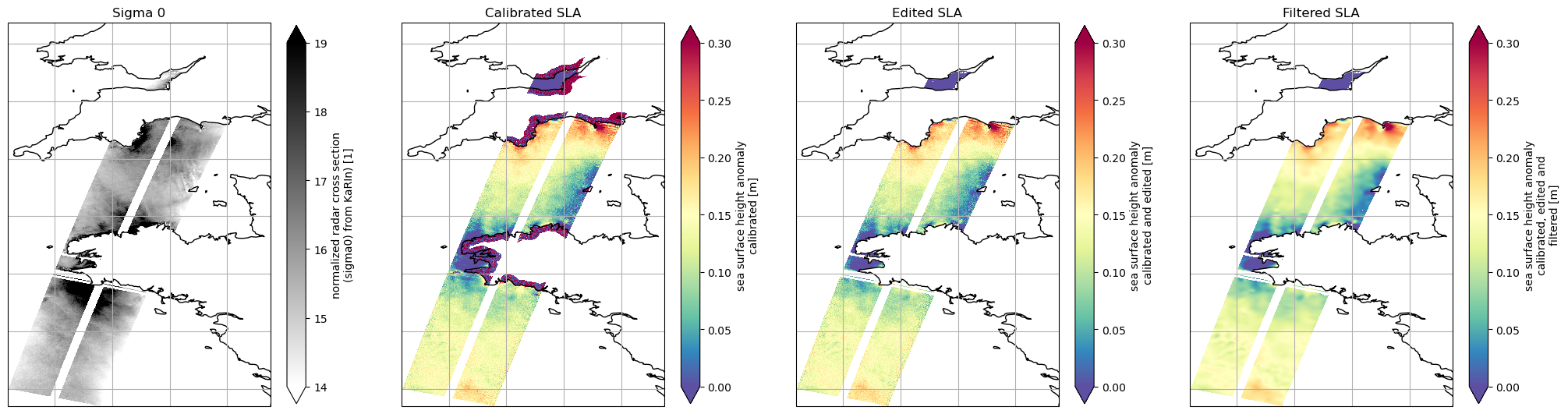
Let’s plot SWOT KaRIn Sigma 0 and the different SLA available
# Apply quality flag
ds_unsmoothed_169["sigma0"] = ds_unsmoothed_169.sigma0.where(ds_unsmoothed_169.quality_flag==0)
# Apply log 10
ds_unsmoothed_169["sigma0_log"] = 10*np.log10(ds_unsmoothed_169["sigma0"])
localbox_zoom = [-5, -1.5, 46, 51]
# set figure
fig, (ax1, ax2, ax3, ax4) = plt.subplots(1, 4, figsize=(25, 21), subplot_kw=dict(projection=ccrs.PlateCarree()))
plot_kwargs = dict(
x="longitude",
y="latitude",
cmap="Spectral_r",
vmin=0,
vmax=0.3,
cbar_kwargs={"shrink": 0.3}
)
plot_kwargs2 = dict(
x="longitude",
y="latitude",
cmap="gray_r",
vmin=14,
vmax=19,
cbar_kwargs={"shrink": 0.3}
)
ds_unsmoothed_169.sigma0_log.plot.pcolormesh(ax=ax1, **plot_kwargs2)
ax1.set_title("Sigma 0")
ds_unsmoothed_169.ssha_unedited.plot.pcolormesh(ax=ax2, **plot_kwargs)
ax2.set_title("Calibrated SLA")
ds_unsmoothed_169.ssha_unfiltered.plot.pcolormesh(ax=ax3, **plot_kwargs)
ax3.set_title("Edited SLA")
ds_unsmoothed_169.ssha_filtered.plot.pcolormesh(ax=ax4, **plot_kwargs)
ax4.set_title("Filtered SLA")
for ax in [ax1, ax2, ax3, ax4]:
# ax.set_extent(localbox_zoom)
ax.coastlines()
ax.gridlines()
Visualise SWOT SLA with L4 CMEMS SLA map
Download L4 CMEMS SLA maps with copernicusmarine client
date_pass_14 = str(ds_unsmoothed_14.time.values[0].astype('datetime64[D]'))
date_pass_169 = str(ds_unsmoothed_169.time.values[0].astype('datetime64[D]'))
# Load xarray dataset
duacs_l4 = copernicusmarine.open_dataset(
dataset_id = "cmems_obs-sl_glo_phy-ssh_nrt_allsat-l4-duacs-0.25deg_P1D",
start_datetime=date_pass_14,
end_datetime=date_pass_169,
minimum_longitude = bbox[0],
maximum_longitude = bbox[2],
minimum_latitude = bbox[1],
maximum_latitude = bbox[3],
variables = ['sla']
)
INFO - 2026-02-27T08:31:11Z - Selected dataset version: "202311"
INFO:copernicusmarine:Selected dataset version: "202311"
INFO - 2026-02-27T08:31:11Z - Selected dataset part: "default"
INFO:copernicusmarine:Selected dataset part: "default"
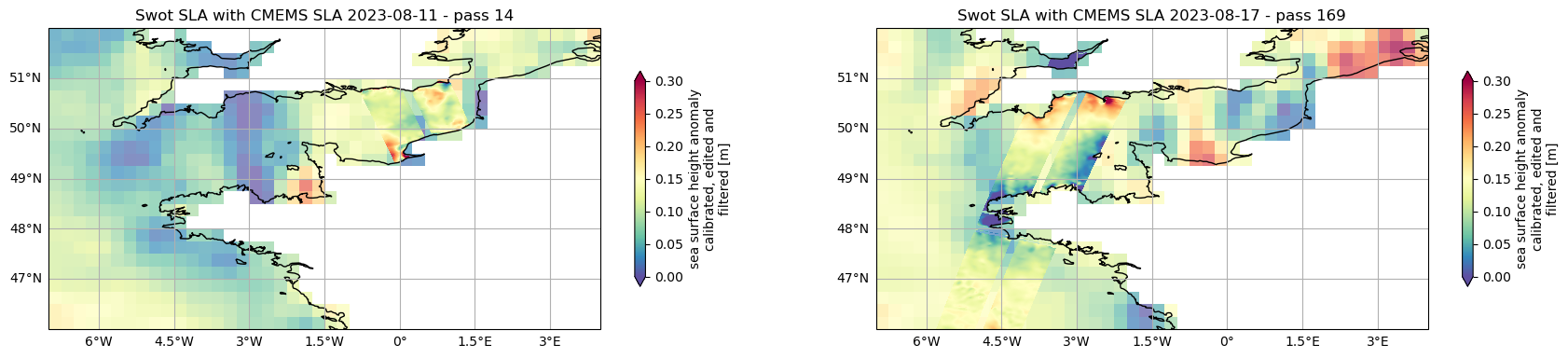
Visualise SWOT SLA with CMEMS SLA
localbox_zoom2 = [-5, -1.5, 46, 51]
plot_kwargs = dict(
x="longitude",
y="latitude",
cmap="Spectral_r",
vmin=0,
vmax=0.3
)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(21, 15), subplot_kw=dict(projection=ccrs.PlateCarree()))
duacs_l4.sla.sel(time=np.datetime64(date_pass_14), method='nearest').plot.pcolormesh(
ax=ax1,
alpha=0.7,
add_colorbar=False,
**plot_kwargs)
ds_unsmoothed_14['ssha_filtered'].plot.pcolormesh(
ax=ax1,
cbar_kwargs={"shrink": 0.2},
**plot_kwargs)
ax1.set_title(f'Swot SLA with CMEMS SLA {date_pass_14} - pass 14')
ax1.set_extent(localbox)
duacs_l4.sla.sel(time=np.datetime64(date_pass_169), method='nearest').plot.pcolormesh(
ax=ax2,
alpha=0.7,
add_colorbar=False,
**plot_kwargs)
ds_unsmoothed_169['ssha_filtered'].plot.pcolormesh(
ax=ax2,
cbar_kwargs={"shrink": 0.2},
**plot_kwargs)
ax2.set_title(f'Swot SLA with CMEMS SLA {date_pass_169} - pass 169')
ax2.set_extent(localbox)
for ax in [ax1, ax2]:
ax.coastlines()
gls = ax.gridlines(draw_labels=True)
gls.top_labels=False
gls.right_labels=False
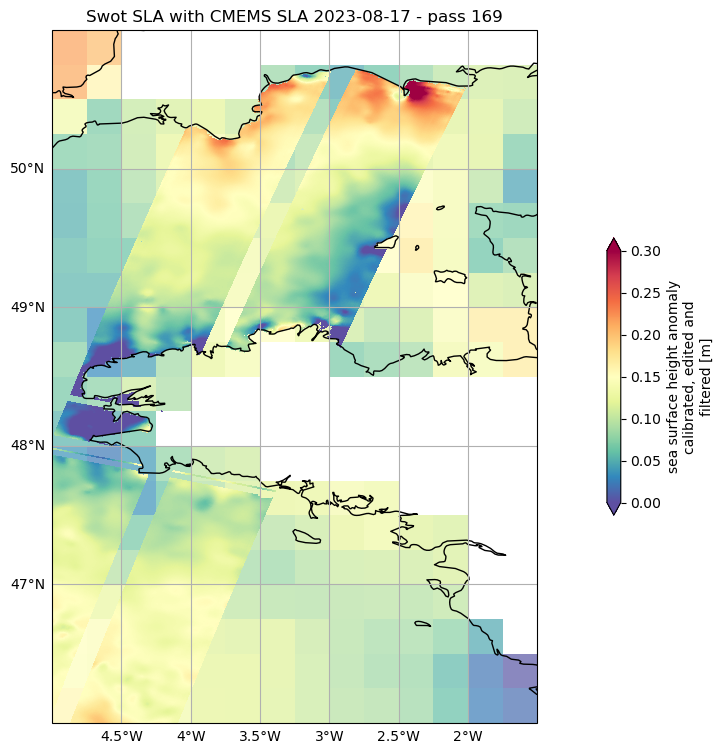
fig, ax = plt.subplots(figsize=(18, 9), subplot_kw=dict(projection=ccrs.PlateCarree()))
duacs_l4.sla.sel(time=np.datetime64(date_pass_169), method='nearest').plot.pcolormesh(
ax=ax,
alpha=0.7,
add_colorbar=False,
**plot_kwargs)
ds_unsmoothed_169['ssha_filtered'].plot.pcolormesh(
ax=ax,
cbar_kwargs={"shrink": 0.4},
**plot_kwargs)
ax.set_title(f'Swot SLA with CMEMS SLA {date_pass_169} - pass 169')
ax.set_extent(localbox_zoom2)
ax.coastlines()
gls = ax.gridlines(draw_labels=True)
gls.top_labels=False
gls.right_labels=False
Visualise SWOT SLA with IFREMER SST map
Download SST maps with copernicusmarine client
# Load xarray dataset
sst = copernicusmarine.open_dataset(
dataset_id = "IFREMER-GLOB-SST-L3-NRT-OBS_FULL_TIME_SERIE",
start_datetime=date_pass_14,
end_datetime=date_pass_169,
minimum_longitude = bbox[0],
maximum_longitude = bbox[2],
minimum_latitude = bbox[1],
maximum_latitude = bbox[3],
variables = ['sea_surface_temperature']
)
INFO - 2026-02-27T08:34:40Z - Selected dataset version: "202211"
INFO:copernicusmarine:Selected dataset version: "202211"
INFO - 2026-02-27T08:34:40Z - Selected dataset part: "default"
INFO:copernicusmarine:Selected dataset part: "default"
ds_sst = sst.sel(time=np.datetime64(date_pass_169), method='nearest')
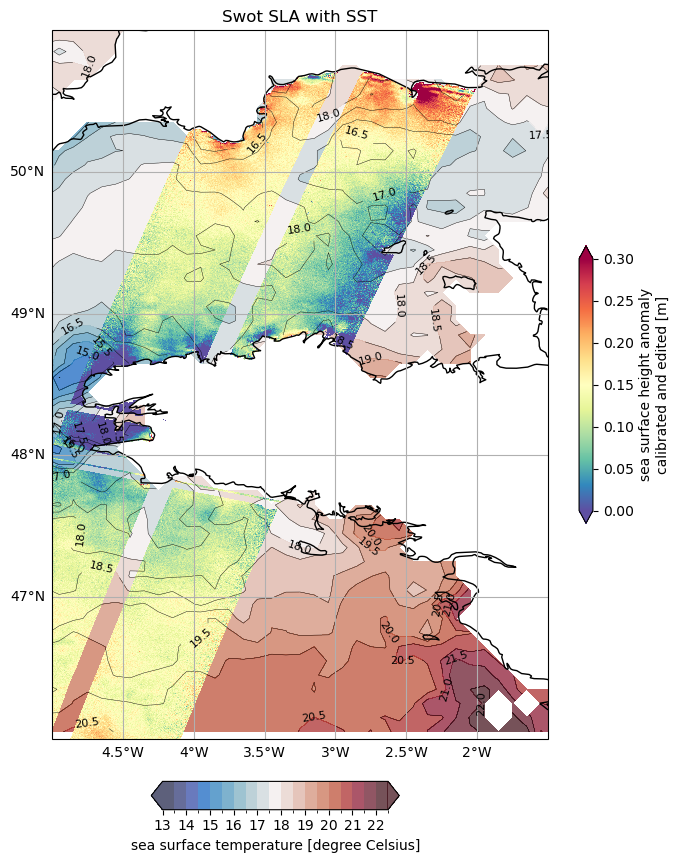
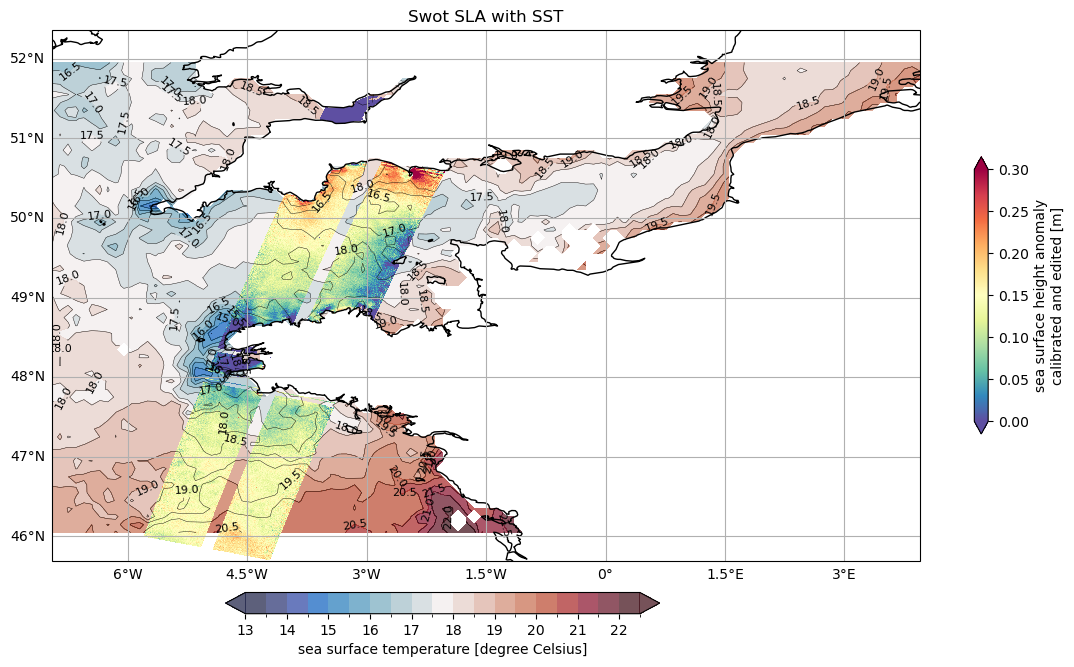
Visualise SWOT SLA with SST
fig, ax = plt.subplots(figsize=(14, 12), subplot_kw=dict(projection=ccrs.PlateCarree()))
plot_kwargs = dict(
x="longitude",
y="latitude",
cmap="Spectral_r"
)
cb_args = dict(
add_colorbar=True,
cbar_kwargs={"shrink": 0.3}
)
cmap_sst = cmocean.cm.balance
levs_sst = np.arange(13,23,step=0.5)
norm_sst = mplcm.colors.BoundaryNorm(levs_sst,cmap_sst.N)
contourf = ax.contourf(ds_sst.longitude,
ds_sst.latitude,
ds_sst.sea_surface_temperature-273.15,
levs_sst,
cmap=cmap_sst,
norm=norm_sst,
alpha=0.7,
extend='both')
contour = ax.contour(ds_sst.longitude,
ds_sst.latitude,
ds_sst.sea_surface_temperature-273.15,
levels=levs_sst,
colors="black",
linewidths=0.3)
ax.clabel(contour, inline=1, fontsize=8)
cax = ax.inset_axes([0.2, -0.1, 0.5, 0.04])
cb_sst = plt.colorbar(contourf, cax=cax, orientation='horizontal', pad=0.005, shrink=0.5)
cb_sst.set_label('sea surface temperature [degree Celsius]')
cb_sst.ax.tick_params(labelsize=10)
ds_unsmoothed_169['ssha_unfiltered'].plot.pcolormesh(
ax=ax,
vmin=0,
vmax=0.3,
**plot_kwargs,
**cb_args)
ax.coastlines()
gls = ax.gridlines(draw_labels=True)
gls.top_labels=False
gls.right_labels=False
ax.set_title('Swot SLA with SST')
Text(0.5, 1.0, 'Swot SLA with SST')
fig, ax = plt.subplots(figsize=(14, 12), subplot_kw=dict(projection=ccrs.PlateCarree()))
contourf = ax.contourf(ds_sst.longitude,
ds_sst.latitude,
ds_sst.sea_surface_temperature-273.15,
levs_sst,
cmap=cmap_sst,
norm=norm_sst,
alpha=0.7,
extend='both')
contour = ax.contour(ds_sst.longitude,
ds_sst.latitude,
ds_sst.sea_surface_temperature-273.15,
levels=levs_sst,
colors="black",
linewidths=0.3)
ax.clabel(contour, inline=1, fontsize=8)
cax = ax.inset_axes([0.2, -0.1, 0.5, 0.04])
cb_sst = plt.colorbar(contourf, cax=cax, orientation='horizontal', pad=0.005, shrink=0.5)
cb_sst.set_label('sea surface temperature [degree Celsius]')
cb_sst.ax.tick_params(labelsize=10)
ds_unsmoothed_169['ssha_unfiltered'].plot.pcolormesh(
ax=ax,
vmin=0,
vmax=0.3,
**plot_kwargs,
**cb_args)
ax.coastlines()
gls = ax.gridlines(draw_labels=True)
gls.top_labels=False
gls.right_labels=False
ax.set_title('Swot SLA with SST')
fig.set_figwidth(8)
ax.set_extent(localbox_zoom2)